Automated Cytometry Data Analysis with Actionable Insights.

terraFlow is a cloud-based platform for automated flow cytometry analysis and literature mapping.

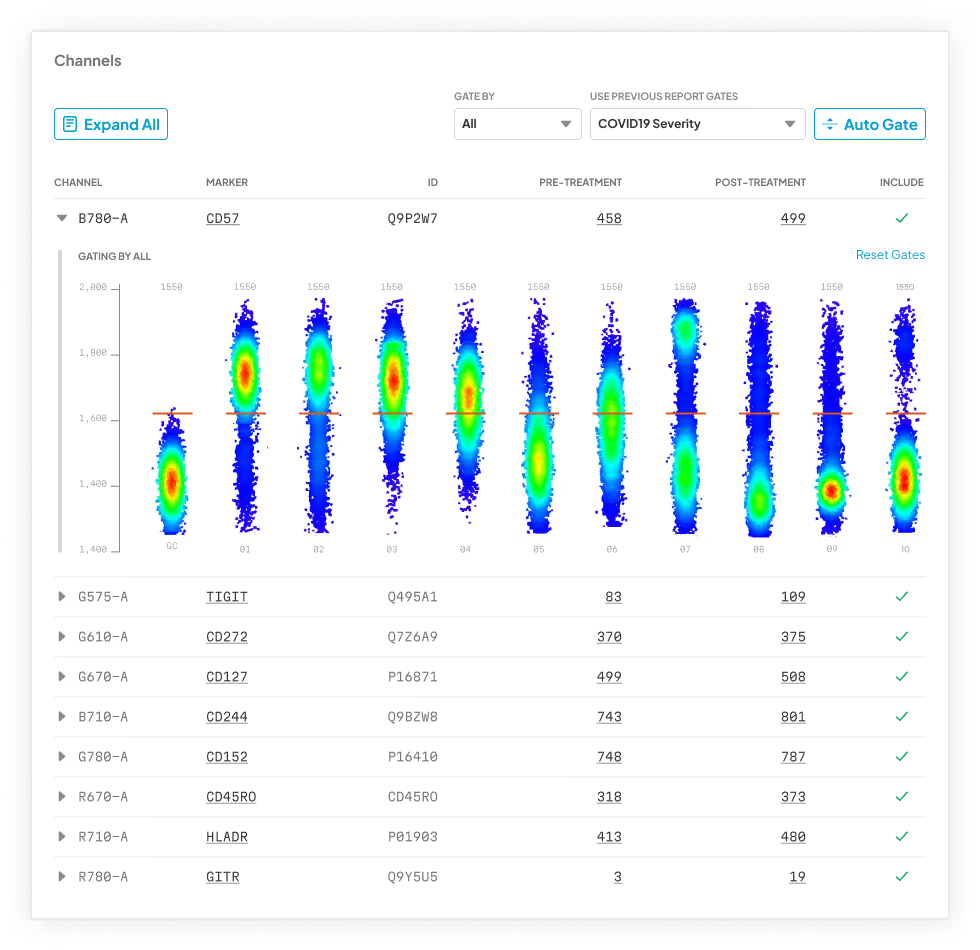
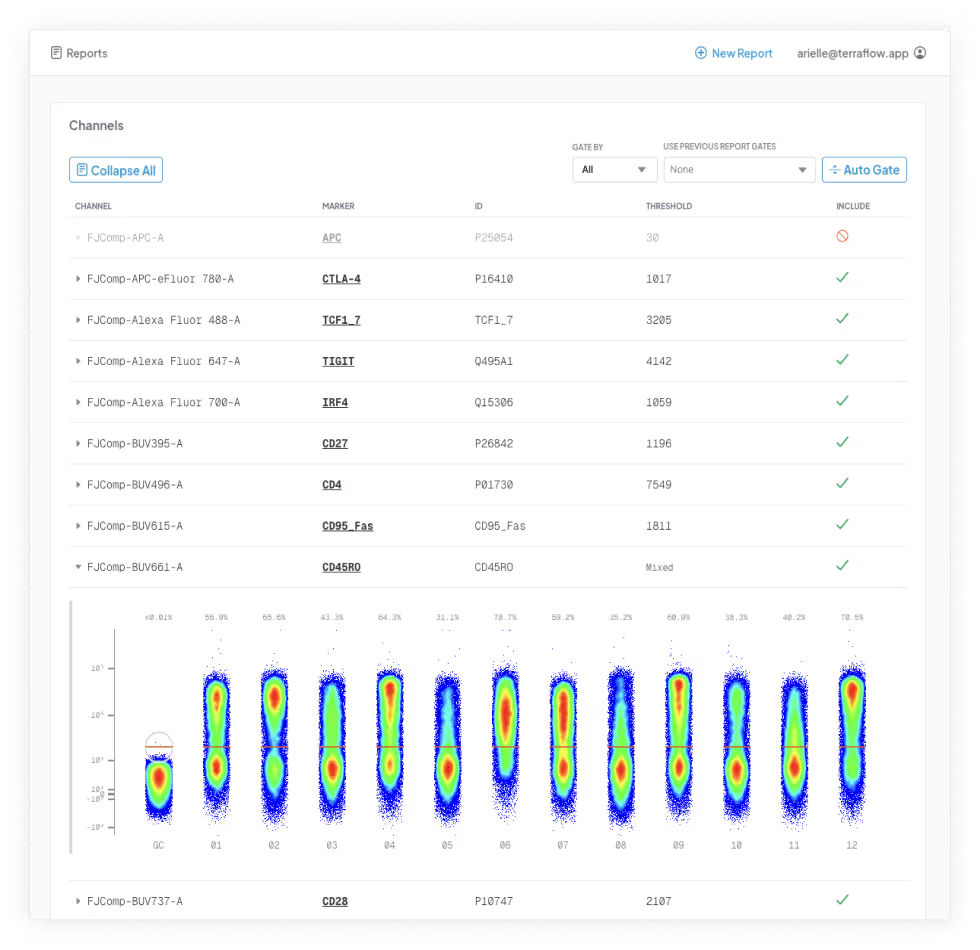
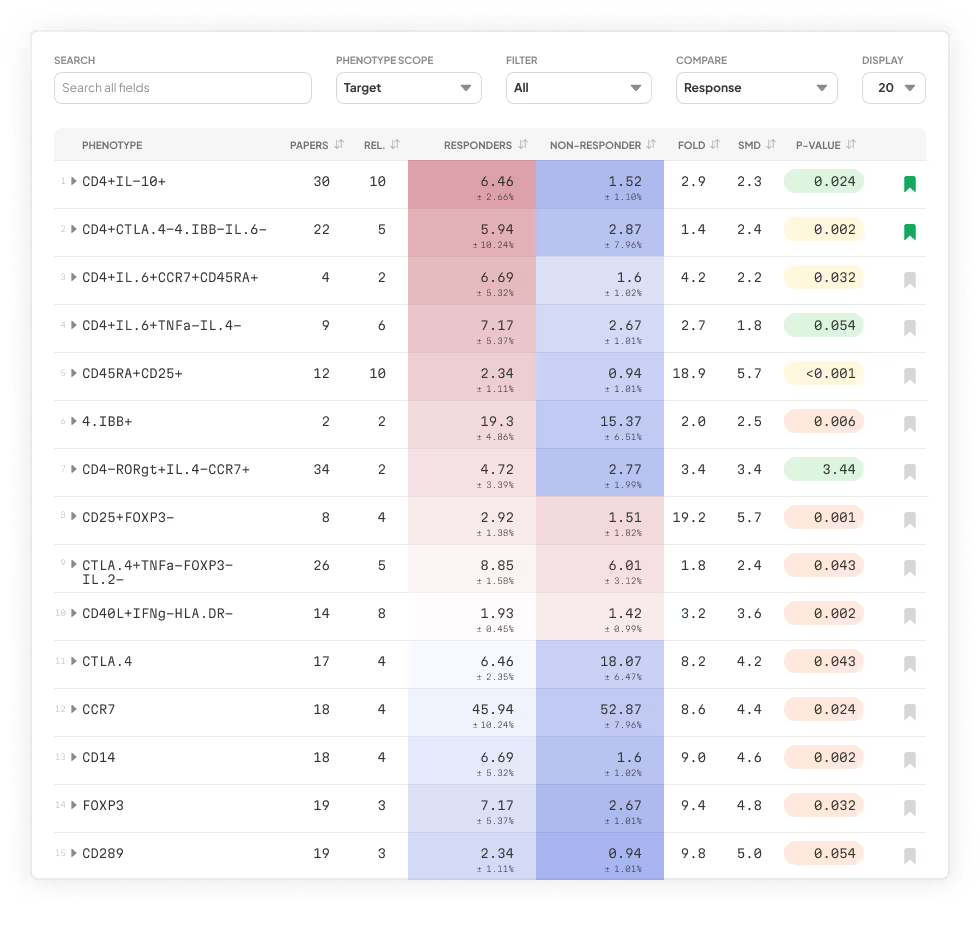
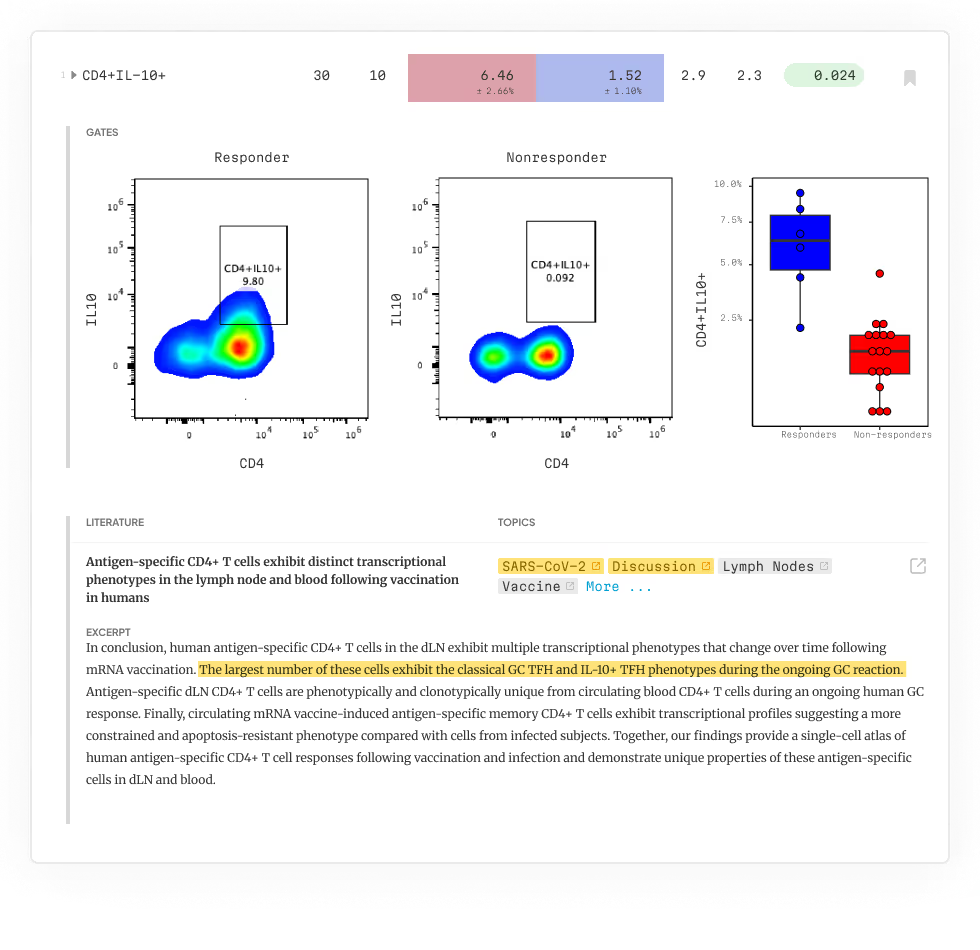
Our platform analyzes every possible phenotype in your cytometry dataset using machine learning and expert-informed models—no coding, software, or bioinformatics team required. Just upload your FCS files and get a detailed, publication-ready report in 24 hours, with clear phenotypes and key findings contextualized and linked directly to primary literature.
Trusted by Industry Leaders
terraFlow in Action
Let us help you find every immune signature in your data and deliver clear, specific, replicable phenotypes—every time.
Book a Demo

We are SOC 2 Type II Certified
- Security: Robust protection against unauthorized access and cyber threats.
- Availability: Reliable uptime and service continuity you can count on.
- Processing Integrity: Accurate, timely, and authorized data processing.
- Confidentiality: Strong safeguards for your sensitive and confidential data.
- Privacy: Responsible handling and protection of your personal information.